- Explore MCP Servers
- molecule-mcp
Molecule Mcp
What is Molecule Mcp
Molecule-MCP is a model-context-protocol server designed for molecules, enabling integration between molecular science tools and Claude AI. It allows Claude to interact with these tools directly, functioning as a co-scientist in molecular modeling.
Use cases
Use cases for Molecule-MCP include educational purposes in molecular biology, research in computational chemistry, and any scenario where collaborative molecular modeling is needed with AI assistance.
How to use
To use Molecule-MCP, first ensure that Claude Desktop is installed and running. Then, configure the ‘claude_desktop_config.json’ file to include the paths to the MCP server and the specific molecular tool servers (e.g., PyMOL, ChimeraX). Finally, install the necessary packages and clone the Molecule-MCP repository to set it up.
Key features
Key features of Molecule-MCP include direct interaction with molecular science tools, the ability for Claude AI to assist in molecule modeling, and support for multiple molecular visualization tools through the Model Context Protocol.
Where to use
undefined
Clients Supporting MCP
The following are the main client software that supports the Model Context Protocol. Click the link to visit the official website for more information.
Overview
What is Molecule Mcp
Molecule-MCP is a model-context-protocol server designed for molecules, enabling integration between molecular science tools and Claude AI. It allows Claude to interact with these tools directly, functioning as a co-scientist in molecular modeling.
Use cases
Use cases for Molecule-MCP include educational purposes in molecular biology, research in computational chemistry, and any scenario where collaborative molecular modeling is needed with AI assistance.
How to use
To use Molecule-MCP, first ensure that Claude Desktop is installed and running. Then, configure the ‘claude_desktop_config.json’ file to include the paths to the MCP server and the specific molecular tool servers (e.g., PyMOL, ChimeraX). Finally, install the necessary packages and clone the Molecule-MCP repository to set it up.
Key features
Key features of Molecule-MCP include direct interaction with molecular science tools, the ability for Claude AI to assist in molecule modeling, and support for multiple molecular visualization tools through the Model Context Protocol.
Where to use
undefined
Clients Supporting MCP
The following are the main client software that supports the Model Context Protocol. Click the link to visit the official website for more information.
Content
Molecule-MCP
Molecule-MCP: A model-context-protocol server for molecules.
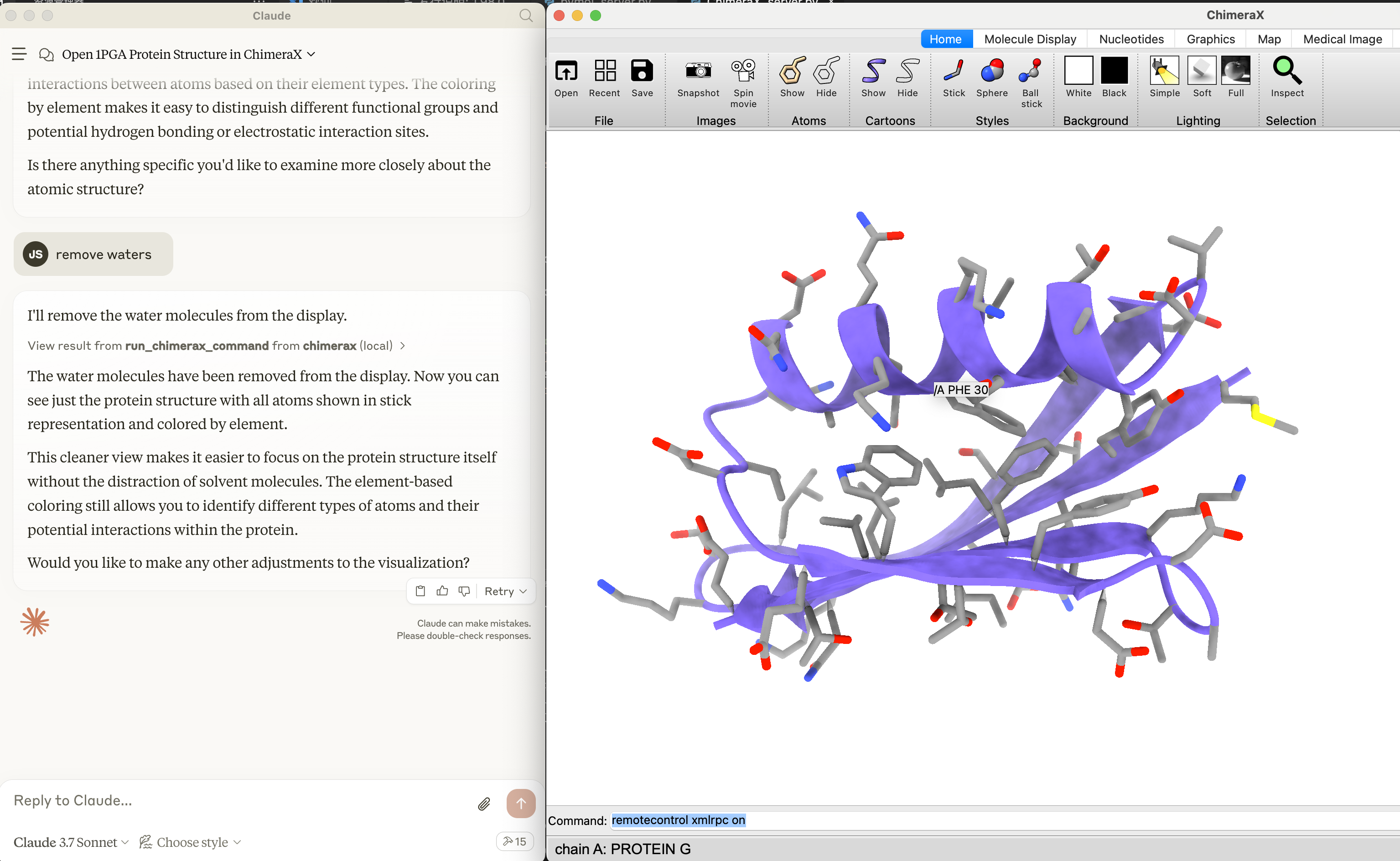
Molecule-MCP connects molecule science releated tools to Claude AI through the Model Context Protocol (MCP), allowing Claude to directly interact with and control these tools and act as a co-scientist. This integration enables prompt assisted molecule modeling.
Installation
⚠️ Note: Molecule-MCP requires Claude Desktop to be installed and running.
- Go to Claude > Settings > Developer > Edit Config > claude_desktop_config.json to include the following:
{
"mcpServers": {
"pymol": {
"command": "/path/to/mcp",
"args": [
"run",
"/path/to/molecule-mcp/pymol_server.py"
]
},
"chimerax": {
"command": "/path/to/mcp",
"args": [
"run",
"/path/to/molecule-mcp/ChimeraX_server.py"
]
},
"gromacs_copilot": {
"command": "/path/to/mcp",
"args": [
"run",
"/path/to/molecule-mcp/mcp_server.py"
]
}
}
}- Install mcp and get the script
pip install "mcp[cli]" chatmol
pip install git+https://github.com/ChatMol/gromacs_copilot.git # optional, for running gromacs_copilot
which mcp
the path to mcp will be displayed. Copy this path for the next step and replace /path/to/mcp with the path to mcp.
git clone https://github.com/ChatMol/molecule-mcp.git cd molecule-mcp pwd
the path to molecule-mcp will be displayed. Copy this path for the next step and replace /path/to/molecule-mcp with the path to molecule-mcp.
Disclaimer
Molecule-MCP is provided “as is” without warranty of any kind, express or implied. The authors and contributors disclaim all warranties including, but not limited to, the implied warranties of merchantability and fitness for a particular purpose. Users employ this software at their own risk.
The authors bear no responsibility for any consequences arising from the use, misuse, or misinterpretation of this software or its outputs. Results obtained through Molecule-MCP should be independently validated prior to use in research, publications, or decision-making processes.
This software is intended for research and educational purposes only. Users are solely responsible for ensuring compliance with applicable laws, regulations, and ethical standards in their jurisdiction.
Dev Tools Supporting MCP
The following are the main code editors that support the Model Context Protocol. Click the link to visit the official website for more information.










