- Explore MCP Servers
- symbolica-mcp
Symbolica Mcp
What is Symbolica Mcp
symbolica-mcp is a scientific computing Model Context Protocol (MCP) server designed for symbolic computation and mathematical calculations, particularly in the field of quantum computing.
Use cases
Use cases for symbolica-mcp include performing complex mathematical calculations, analyzing scientific data, visualizing results, and integrating with AI applications for enhanced computational capabilities.
How to use
To use symbolica-mcp, you can pull the Docker image from Docker Hub and run it in a containerized environment. For integration with Claude for Desktop, specific configuration settings must be added based on your operating system.
Key features
Key features include support for scientific computing libraries like NumPy and SciPy, symbolic mathematics, differential equation solving, linear algebra operations, quantum computing analysis, data visualization with Matplotlib, machine learning with scikit-learn, and cross-platform compatibility.
Where to use
symbolica-mcp is applicable in scientific and engineering fields, especially for tasks involving quantum computing, data analysis, and machine learning.
Clients Supporting MCP
The following are the main client software that supports the Model Context Protocol. Click the link to visit the official website for more information.
Overview
What is Symbolica Mcp
symbolica-mcp is a scientific computing Model Context Protocol (MCP) server designed for symbolic computation and mathematical calculations, particularly in the field of quantum computing.
Use cases
Use cases for symbolica-mcp include performing complex mathematical calculations, analyzing scientific data, visualizing results, and integrating with AI applications for enhanced computational capabilities.
How to use
To use symbolica-mcp, you can pull the Docker image from Docker Hub and run it in a containerized environment. For integration with Claude for Desktop, specific configuration settings must be added based on your operating system.
Key features
Key features include support for scientific computing libraries like NumPy and SciPy, symbolic mathematics, differential equation solving, linear algebra operations, quantum computing analysis, data visualization with Matplotlib, machine learning with scikit-learn, and cross-platform compatibility.
Where to use
symbolica-mcp is applicable in scientific and engineering fields, especially for tasks involving quantum computing, data analysis, and machine learning.
Clients Supporting MCP
The following are the main client software that supports the Model Context Protocol. Click the link to visit the official website for more information.
Content
symbolica-mcp
A scientific computing Model Context Protocol (MCP) server allows AI, such as Claude, to perform symbolic computing, conduct calculations, analyze data, and generate visualizations. This is particularly useful for scientific and engineering applications, including quantum computing, all within a containerized environment.
Features
- Run scientific computing operations with NumPy, SciPy, SymPy, Pandas
- Perform symbolic mathematics and solve differential equations
- Support for linear algebra operations and matrix manipulations
- Quantum computing analysis
- Create data visualizations with Matplotlib and Seaborn
- Perform machine learning operations with scikit-learn
- Execute tensor operations and complex matrix calculations
- Analyze data sets with statistical tools
- Cross-platform support (automatically detects Windows, macOS, and Linux), especially for users with Mac M series chips
- Works on both Intel/AMD (x86_64) and ARM processors
Quick Start
Using the Docker image
# Pull the image from Docker Hub
docker pull ychen94/computing-mcp:latest
# Run the container (automatically detects host OS)
docker run -i --rm -v /tmp:/app/shared ychen94/computing-mcp:latest
For Windows users:
docker run -i --rm -v $env:TEMP:/app/shared ychen94/computing-mcp:latest
Integrating with Claude for Desktop
- Open Claude for Desktop
- Open Settings ➝ Developer ➝ Edit Config
- Add the following configuration:
For MacOS/Linux:
{
"mcpServers": {
"computing-mcp": {
"command": "docker",
"args": [
"run",
"-i",
"--rm",
"-v",
"/tmp:/app/shared",
"ychen94/computing-mcp:latest"
]
}
}
}For Windows:
{
"mcpServers": {
"computing-mcp": {
"command": "docker",
"args": [
"run",
"-i",
"--rm",
"-v",
"%TEMP%:/app/shared",
"ychen94/computing-mcp:latest"
]
}
}
}Examples
Tensor Products
Can you calculate and visualize the tensor product of two matrices? Please run: import numpy as np import matplotlib.pyplot as plt # Define two matrices A = np.array([[1, 2], [3, 4]]) B = np.array([[5, 6], [7, 8]]) # Calculate tensor product using np.kron (Kronecker product) tensor_product = np.kron(A, B) # Display the result print("Matrix A:") print(A) print("\nMatrix B:") print(B) print("\nTensor Product A ⊗ B:") print(tensor_product) # Create a visualization of the tensor product plt.figure(figsize=(8, 6)) plt.imshow(tensor_product, cmap='viridis') plt.colorbar(label='Value') plt.title('Visualization of Tensor Product A ⊗ B')
Symbolic Mathematics
Can you solve this differential equation? Please run: import sympy as sp import matplotlib.pyplot as plt import numpy as np # Define symbolic variable x = sp.Symbol('x') y = sp.Function('y')(x) # Define the differential equation: y''(x) + 2*y'(x) + y(x) = 0 diff_eq = sp.Eq(sp.diff(y, x, 2) + 2*sp.diff(y, x) + y, 0) # Solve the equation solution = sp.dsolve(diff_eq) print("Solution:") print(solution) # Plot a particular solution (C1=1, C2=0) solution_func = solution.rhs.subs({sp.symbols('C1'): 1, sp.symbols('C2'): 0}) print("Particular solution:") print(solution_func) # Create a numerical function we can evaluate solution_lambda = sp.lambdify(x, solution_func) # Plot the solution x_vals = np.linspace(0, 5, 100) y_vals = [float(solution_lambda(x_val)) for x_val in x_vals] plt.figure(figsize=(10, 6)) plt.plot(x_vals, y_vals) plt.grid(True) plt.title("Solution to y''(x) + 2*y'(x) + y(x) = 0") plt.xlabel('x') plt.ylabel('y(x)') plt.show()
Data Analysis
Can you perform a clustering analysis on this dataset? Please run: import numpy as np import pandas as pd import matplotlib.pyplot as plt from sklearn.cluster import KMeans from sklearn.preprocessing import StandardScaler # Create a sample dataset np.random.seed(42) n_samples = 300 # Create three clusters cluster1 = np.random.normal(loc=[2, 2], scale=0.5, size=(n_samples//3, 2)) cluster2 = np.random.normal(loc=[7, 7], scale=0.5, size=(n_samples//3, 2)) cluster3 = np.random.normal(loc=[2, 7], scale=0.5, size=(n_samples//3, 2)) # Combine clusters X = np.vstack([cluster1, cluster2, cluster3]) # Create DataFrame df = pd.DataFrame(X, columns=['Feature1', 'Feature2']) print(df.head()) # Standardize data scaler = StandardScaler() X_scaled = scaler.fit_transform(X) # Apply KMeans clustering kmeans = KMeans(n_clusters=3, random_state=42) df['Cluster'] = kmeans.fit_predict(X_scaled) # Plot the clusters plt.figure(figsize=(10, 6)) for cluster_id in range(3): cluster_data = df[df['Cluster'] == cluster_id] plt.scatter(cluster_data['Feature1'], cluster_data['Feature2'], label=f'Cluster {cluster_id}', alpha=0.7) # Plot cluster centers centers = scaler.inverse_transform(kmeans.cluster_centers_) plt.scatter(centers[:, 0], centers[:, 1], s=200, c='red', marker='X', label='Centers') plt.title('K-Means Clustering Results') plt.xlabel('Feature 1') plt.ylabel('Feature 2') plt.legend() plt.grid(True)
Quantum Computing

Gallery
laser physics:

elliptic integral:
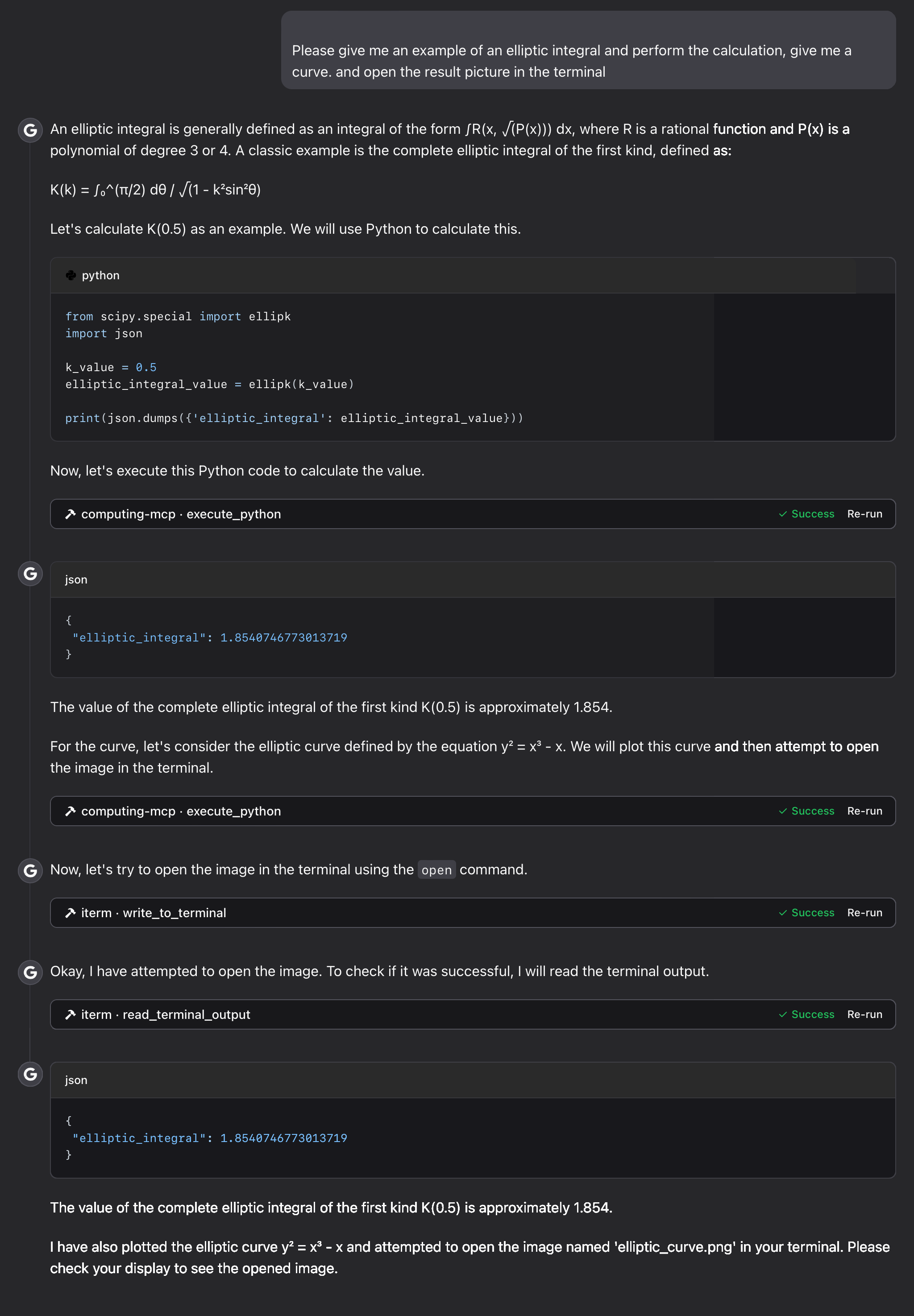
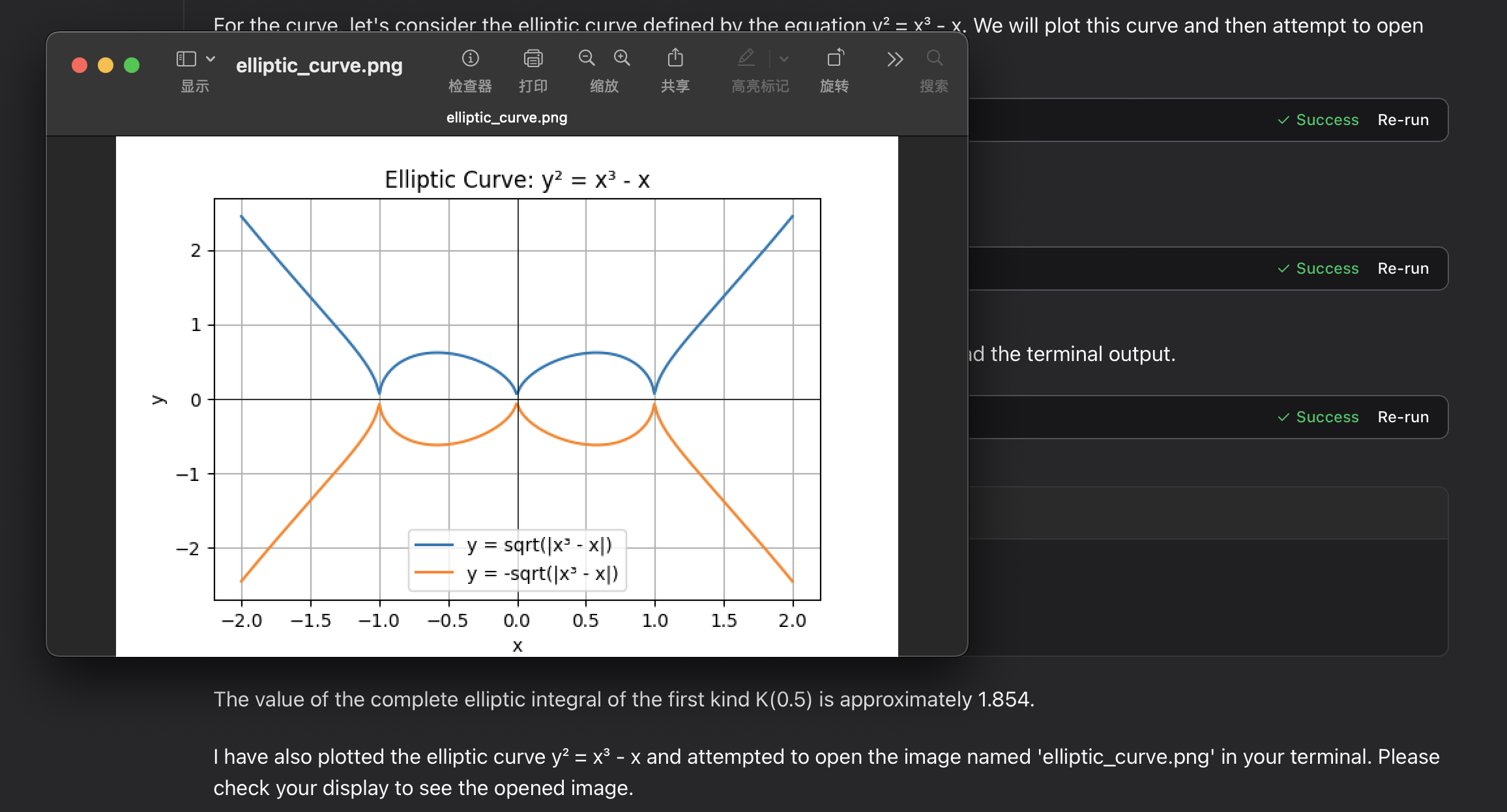
Troubleshooting
Common Issues
-
Permission errors with volume mounts
- Ensure the mount directory exists and has appropriate permissions
-
Plot pciture files not appearing
-
Check the path in your host system:
/tmpfor macOS/Linux or your temp folder for Windows -
Verify Docker has permissions to write to the mount location
-
check the mcp tool’s output content
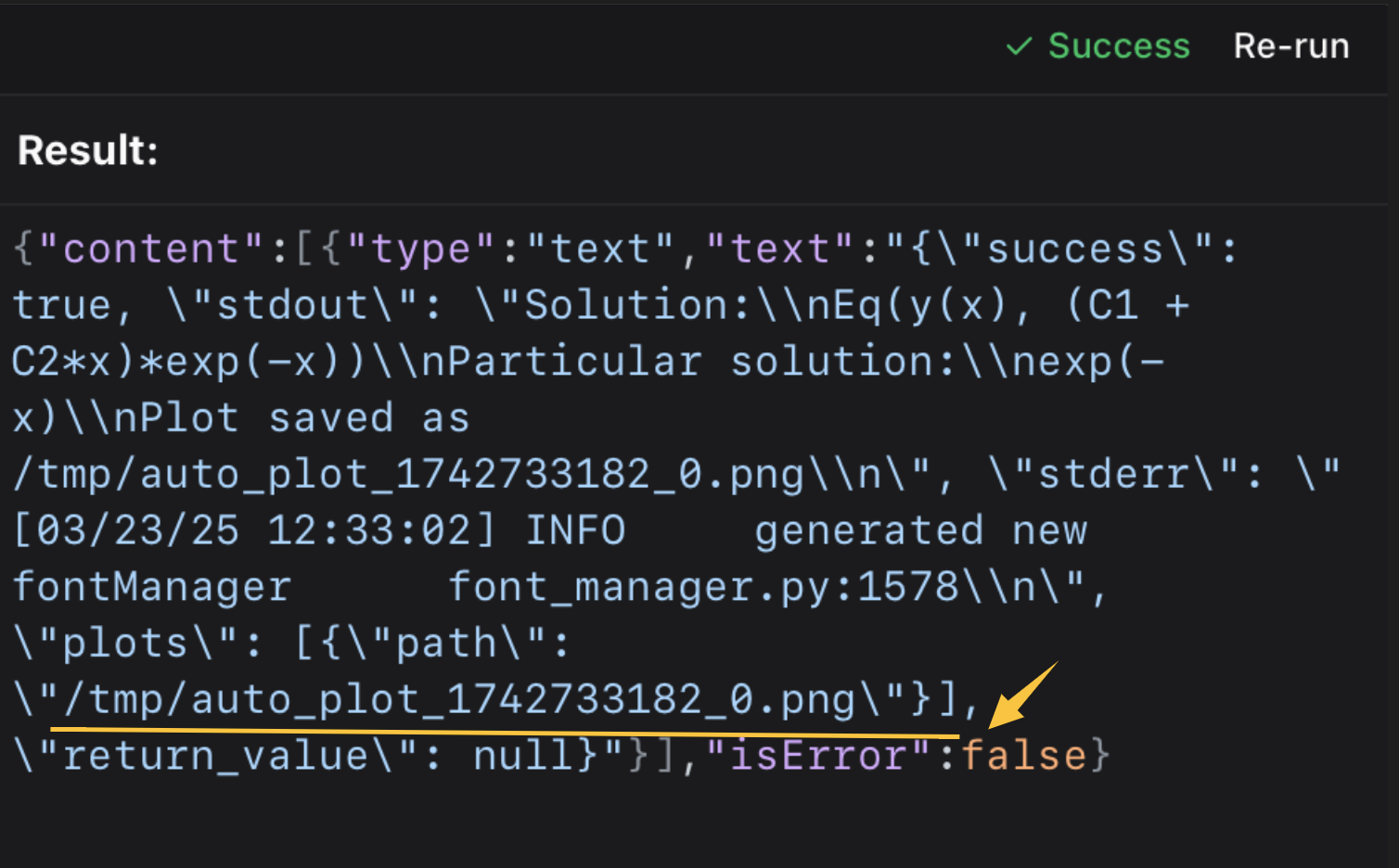
then open it in the terminal or your picture viewer.⭐️ ⭐️
I use the iterm-mcp-server or other terminals’ mcp servers to open the file without interrupting your workflow.
⭐️ ⭐️
-
Support
If you encounter issues, please open a GitHub issue with:
- Error messages
- Your operating system and Docker version
- Steps to reproduce the problem
License
This project is licensed under the MIT License.
For more details, please see the LICENSE file in this project repository.
Dev Tools Supporting MCP
The following are the main code editors that support the Model Context Protocol. Click the link to visit the official website for more information.










